plot2 is a high-level R wrapper around ggplot2 designed to minimise typing while producing complex, publication-ready plots. The central idea: one call to plot2() replaces many separate ggplot2 + dplyr/tidyr steps, including automatic plot-type selection, in-line data transformations, axis handling, and custom theming.
head(iris)
#> Sepal.Length Sepal.Width Petal.Length Petal.Width Species
#> 1 5.1 3.5 1.4 0.2 setosa
#> 2 4.9 3.0 1.4 0.2 setosa
#> 3 4.7 3.2 1.3 0.2 setosa
#> 4 4.6 3.1 1.5 0.2 setosa
#> 5 5.0 3.6 1.4 0.2 setosa
#> 6 5.4 3.9 1.7 0.4 setosa
library(ggplot2)
ggplot(
data = iris,
mapping = aes(
x = Species,
# transformation in ggplot do not account for other aesthetics:
y = mean(Sepal.Length))) +
geom_col(width = 0.5) +
scale_y_continuous(expand = expansion(mult = c(0, 0.25)))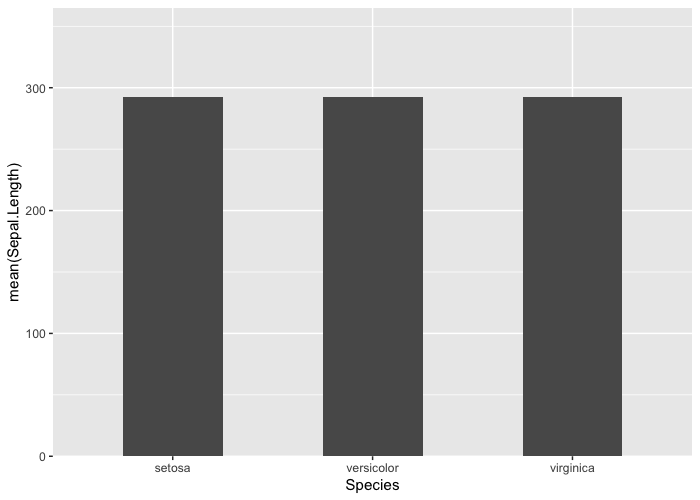
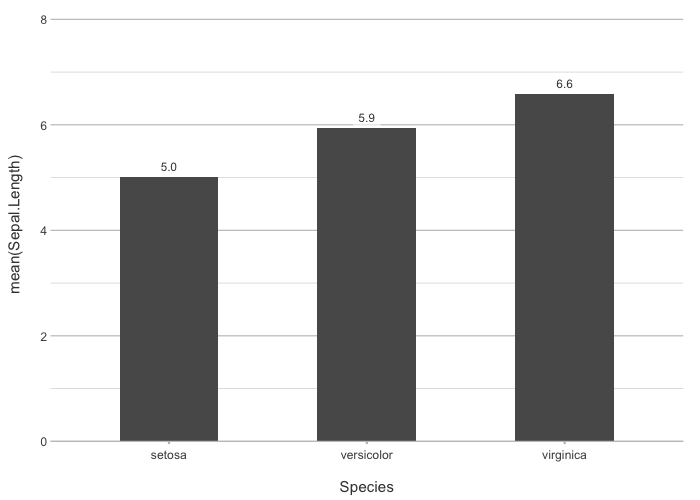
plot2 is designed to streamline the process of creating high-quality data visualisations in R by removing most of the manual work. Built with the philosophy of Less Typing, More Plotting, plot2 automates many of the routine tasks that typically require substantial code when plotting with ggplot2. It even renders pre-processing steps in, for example, dplyr, tidyr, and forcats largely superfluous. This package allows you to focus on the insights and stories your data can tell, rather than the intricate details of plot construction.
Where ggplot2 involves calling many different functions such as ggplot(), aes(), geom_col(), facet_wrap(), theme(), and scale_y_continuous(), plot2 only requires filling in the necessary arguments of a single function.
For a comprehensive guide to using
plot2, including advanced features and customisation options, please see the full vignette here.
Key Features
-
Plotting with as Few Lines as Possible: No need to type
ggplot(),aes(),geom_col(),facet_wrap(),theme(), orscale_y_continuous(). A singleplot2()call is sufficient. -
Automatic Plot Selection:
plot2automatically chooses the most appropriate plot type based on your data, saving time and effort. -
In-line Data Transformations: Eliminate manual pre-processing steps, for example in
dplyrandtidyr, by performing data transformations directly within the plotting function, including axis and plot titles. - Enhanced Sorting and Faceting: Easily sort and facet your data with simple arguments, streamlining the creation of complex multi-panel plots.
-
New Clean Theme: Includes
theme_minimal2(), a minimalist theme based ontheme_minimal(), further optimised for clear and professional output, making it well suited to PDF publications, scientific manuscripts, and presentations. -
Seamless Integration with ggplot2: Retain the full power and flexibility of
ggplot2while benefiting fromplot2’s streamlined interface.
Philosophy
ggplot2 is a versatile tool that has become a cornerstone of data visualisation in R, providing users with substantial control over their plots. However, this flexibility often results in repetitive and verbose code, particularly for routine tasks such as setting axis labels, choosing plot types, or transforming data.
plot2 complements ggplot2 by offering a more streamlined and intuitive interface. It simplifies plot creation by automatically handling many implementation details, without sacrificing flexibility or expressive power. Whether you are exploring data or preparing a publication-ready visualisation, plot2 enables you to work more efficiently with less code.
Installation
Install the latest version of plot2 as follows:
install.packages("plot2",
repos = c("https://cran.r-project.org",
"https://msberends.r-universe.dev"))
# OR:
remotes::install_github("msberends/plot2")Examples
For all supported plot types, see the overview: https://msberends.github.io/plot2/articles/supported_types.html.
Here is a minimal example to get started with plot2:
library(plot2)
head(iris)
#> Sepal.Length Sepal.Width Petal.Length Petal.Width Species
#> 1 5.1 3.5 1.4 0.2 setosa
#> 2 4.9 3.0 1.4 0.2 setosa
#> 3 4.7 3.2 1.3 0.2 setosa
#> 4 4.6 3.1 1.5 0.2 setosa
#> 5 5.0 3.6 1.4 0.2 setosa
#> 6 5.4 3.9 1.7 0.4 setosa
# No variables specified, so plot2() determines them automatically.
# The default type will be points because the first two variables are numeric.
iris |>
plot2()
#> ℹ Using category = Species
#> ℹ Using type = "point" since both axes are numeric
#> ℹ Using x = Sepal.Length
#> ℹ Using y = Sepal.Width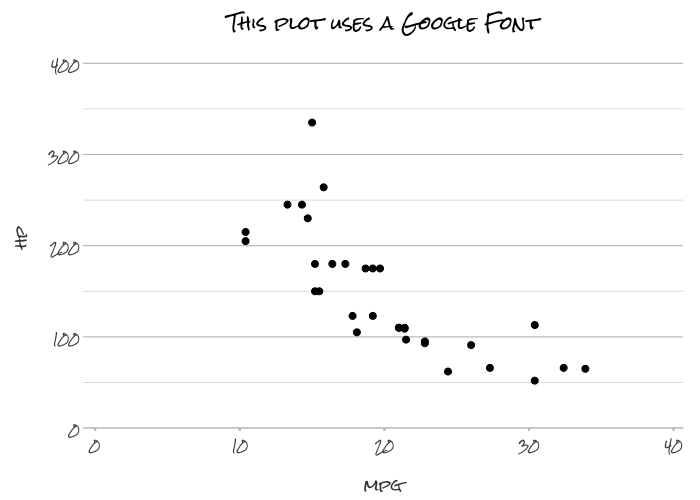
# If x and y are specified, no additional mapping is set:
iris |>
plot2(Sepal.Width, Sepal.Length)
#> ℹ Using type = "point" since both axes are numeric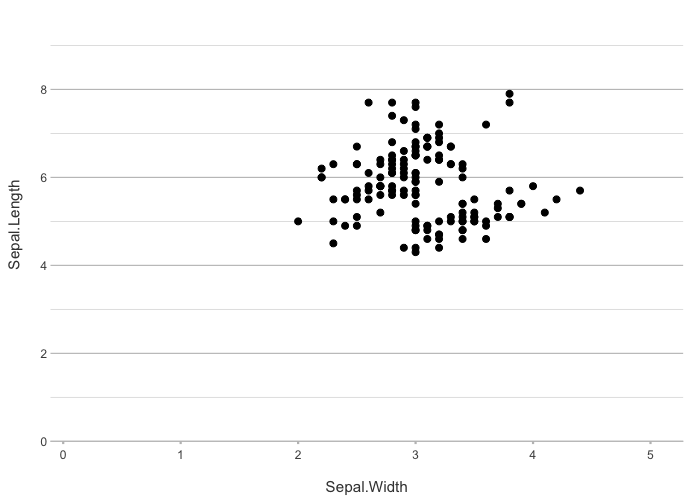
iris |>
plot2(Species, Sepal.Length)
#> ℹ Using type = "boxplot" since all groups in Species contain at least three values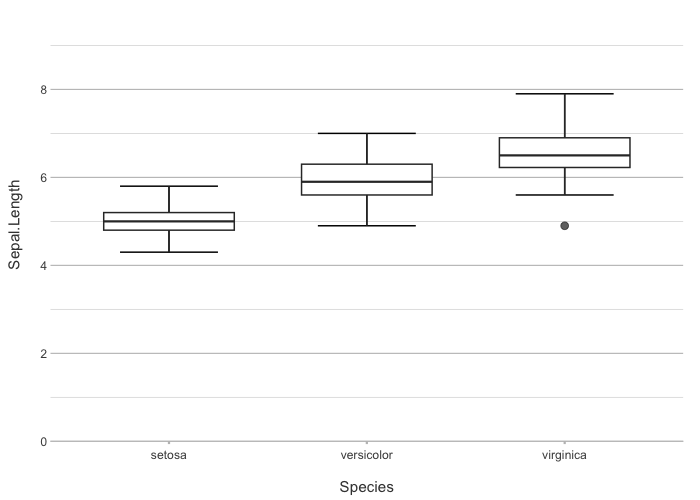
# Arguments are in the order: x, y, category, facet
iris |>
plot2(Sepal.Length, Sepal.Width, Petal.Length, Species)
#> ℹ Assuming facet.fixed_x = TRUE since the three x scales differ by less than 25%
#> ℹ Assuming facet.fixed_y = TRUE since the three y scales differ by less than 25%
#> ℹ Assuming facet.repeat_lbls_y = FALSE since y has fixed scales
#> ℹ Using type = "point" since both axes are numeric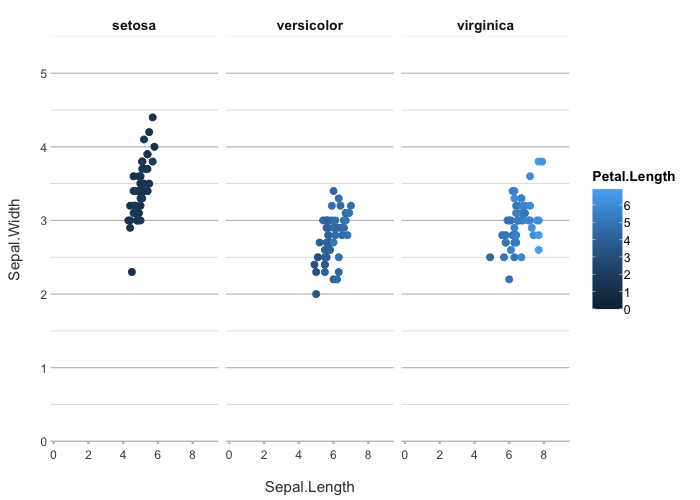
iris |>
plot2(Sepal.Length, Sepal.Width, Petal.Length, Species,
colour = "viridis")
#> ℹ Assuming facet.fixed_x = TRUE since the three x scales differ by less than 25%
#> ℹ Assuming facet.fixed_y = TRUE since the three y scales differ by less than 25%
#> ℹ Assuming facet.repeat_lbls_y = FALSE since y has fixed scales
#> ℹ Using type = "point" since both axes are numeric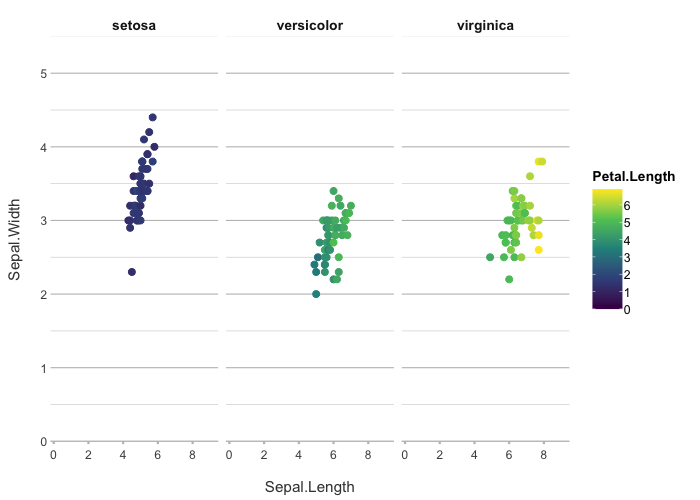
iris |>
plot2(Sepal.Length, Sepal.Width, Petal.Length, Species,
colour = c("white", "red", "black"))
#> ℹ Assuming facet.fixed_x = TRUE since the three x scales differ by less than 25%
#> ℹ Assuming facet.fixed_y = TRUE since the three y scales differ by less than 25%
#> ℹ Assuming facet.repeat_lbls_y = FALSE since y has fixed scales
#> ℹ Using category.midpoint = 3.45 (the current category scale centre)
#> ℹ Using type = "point" since both axes are numeric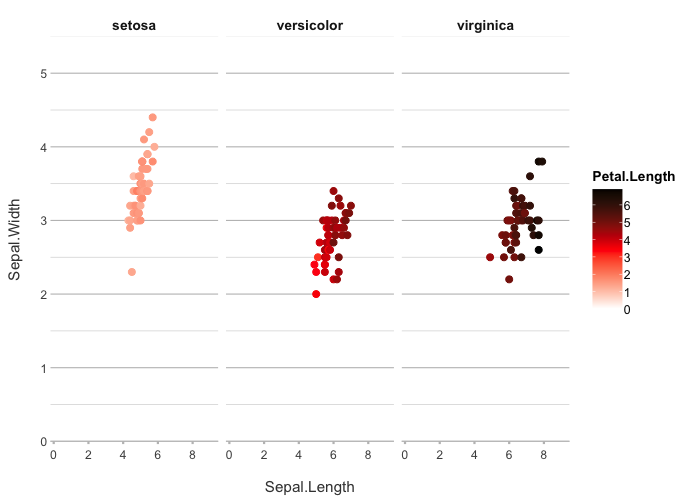
# y can contain multiple named columns
iris |>
plot2(x = Sepal.Length,
y = c(Length = Petal.Length, Width = Petal.Width),
category.title = "Petal property")
#> ℹ Using type = "point" since both axes are numeric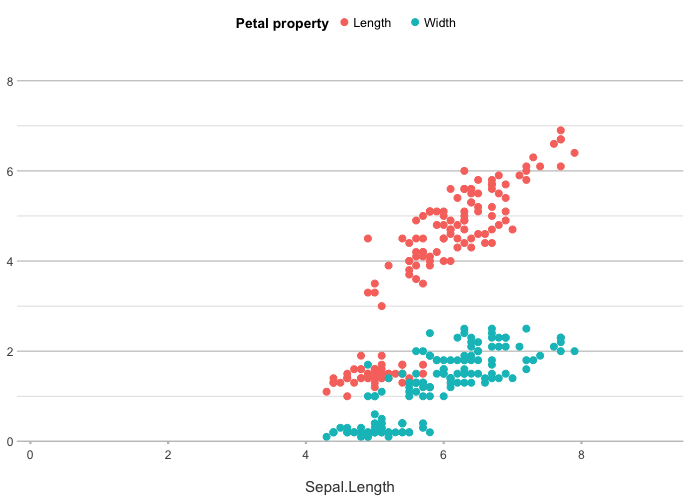
iris |>
plot2(x = Species, y = where(is.double))
#> ℹ Using type = "boxplot" since all groups in Species and category contain at least three values
#> ℹ Using y = c(Petal.Length, Petal.Width, Sepal.Length, Sepal.Width)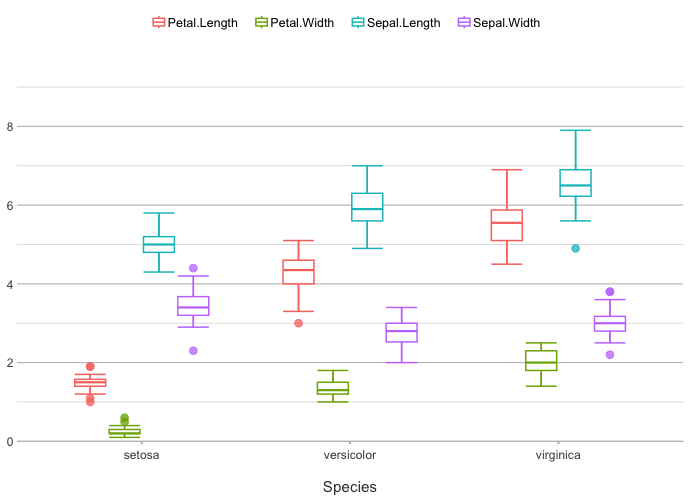
# The category type can consist of one or more aesthetics
iris |>
plot2(zoom = TRUE,
category.type = c("colour", "shape"),
size = 3)
#> ℹ Using category = Species
#> ℹ Using type = "point" since both axes are numeric
#> ℹ Using x = Sepal.Length
#> ℹ Using y = Sepal.Width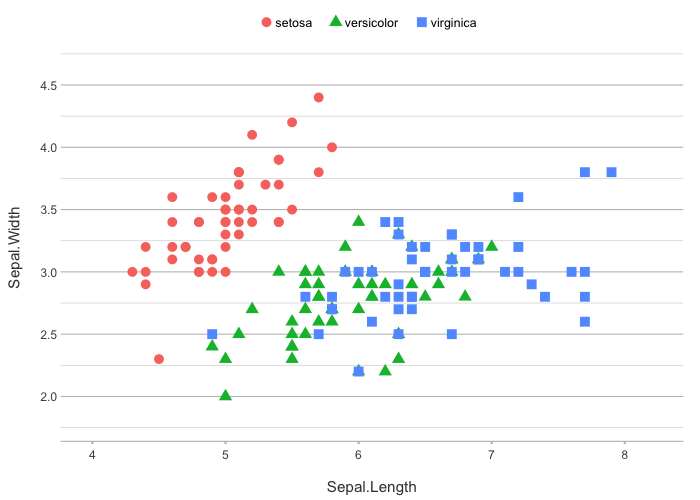
iris |>
plot2(zoom = TRUE,
category = Petal.Length,
category.type = c("colour", "size"),
colour = "viridis")
#> ℹ Using type = "point" since both axes are numeric
#> ℹ Using x = Sepal.Length
#> ℹ Using y = Sepal.Width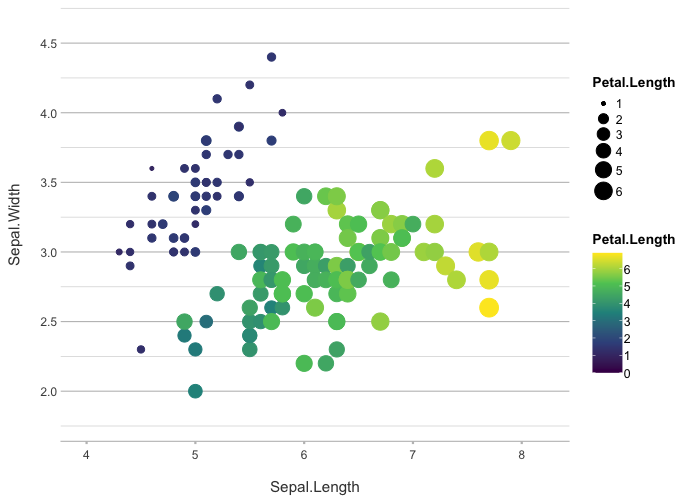
# Easily add smoothing
iris |>
plot2(zoom = TRUE,
smooth = TRUE)
#> ℹ Using category = Species
#> ℹ Using type = "point" since both axes are numeric
#> ℹ Using x = Sepal.Length
#> ℹ Using y = Sepal.Width
#> `geom_smooth()` using method = 'loess' and formula = 'y ~ x'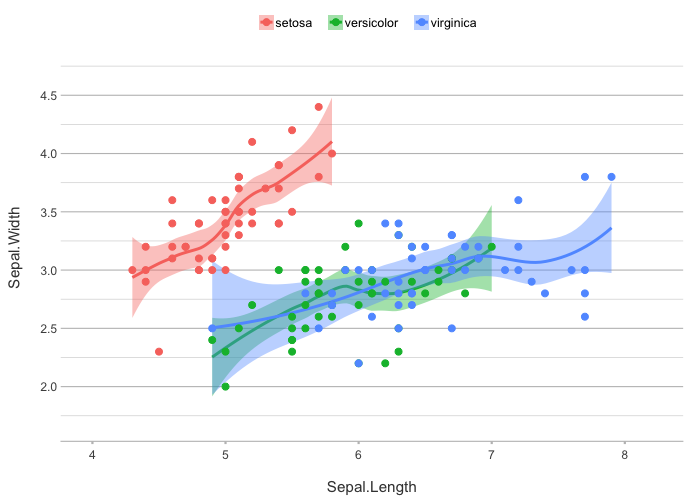
iris |>
plot2(zoom = TRUE,
smooth = TRUE,
smooth.method = "lm")
#> ℹ Using category = Species
#> ℹ Using type = "point" since both axes are numeric
#> ℹ Using x = Sepal.Length
#> ℹ Using y = Sepal.Width
#> `geom_smooth()` using formula = 'y ~ x'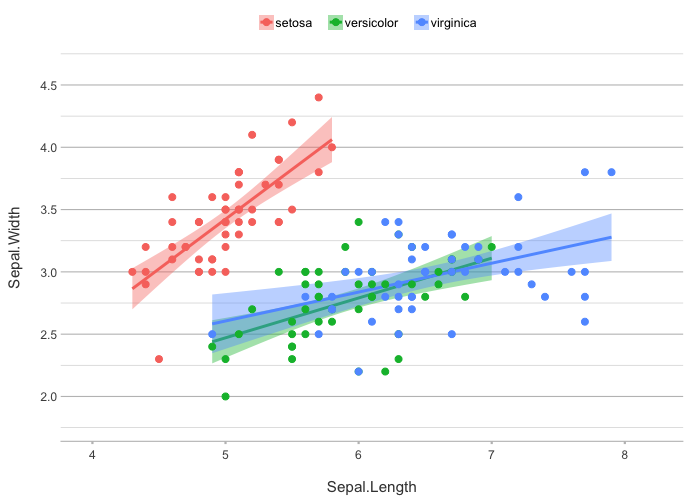
# Support for a secondary y-axis
mtcars |>
plot2(x = mpg,
y = hp,
y_secondary = disp ^ 2,
y_secondary.scientific = TRUE,
title = "Secondary y-axis sets colour to the axis titles")
#> ℹ Using type = "point" since both axes are numeric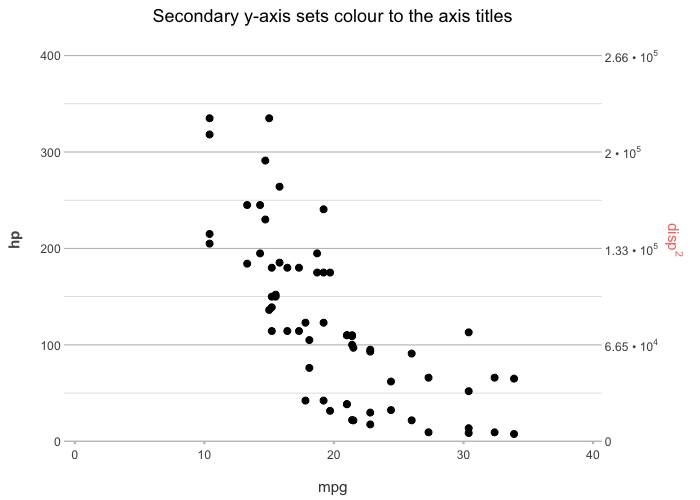
admitted_patients
#> # A tibble: 250 × 7
#> date patient_id gender age age_group hospital ward
#> <date> <dbl> <chr> <dbl> <ord> <fct> <chr>
#> 1 2002-01-14 1 M 78 75+ D Non-ICU
#> 2 2002-03-17 2 M 78 75+ C Non-ICU
#> 3 2002-04-08 3 M 78 75+ A ICU
#> 4 2002-04-14 4 M 72 55-74 C Non-ICU
#> 5 2002-05-07 5 M 83 75+ C Non-ICU
#> 6 2002-05-16 6 F 65 55-74 B ICU
#> 7 2002-05-16 7 M 47 25-54 D Non-ICU
#> 8 2002-06-18 8 M 30 25-54 B ICU
#> 9 2002-06-23 9 M 82 75+ D Non-ICU
#> 10 2002-06-23 9 M 82 75+ D Non-ICU
#> # ℹ 240 more rows
# Arguments are in the order: x, y, category, facet
admitted_patients |>
plot2(hospital, age)
#> ℹ Using type = "boxplot" since all groups in hospital contain at least three values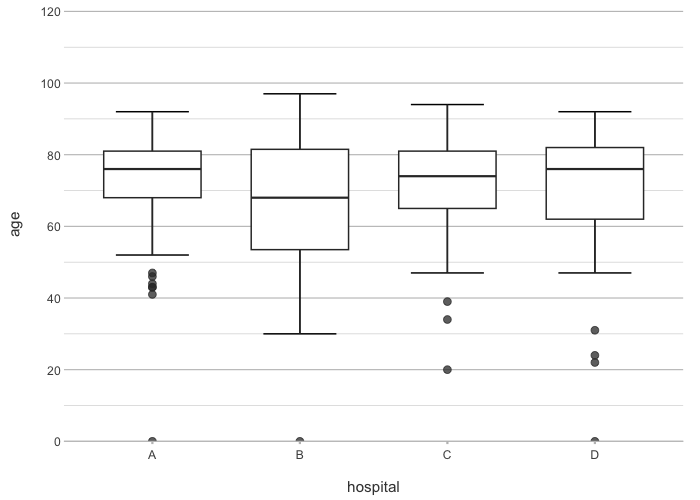
admitted_patients |>
plot2(hospital, age, gender)
#> ℹ Using type = "boxplot" since all groups in hospital and gender contain at least three values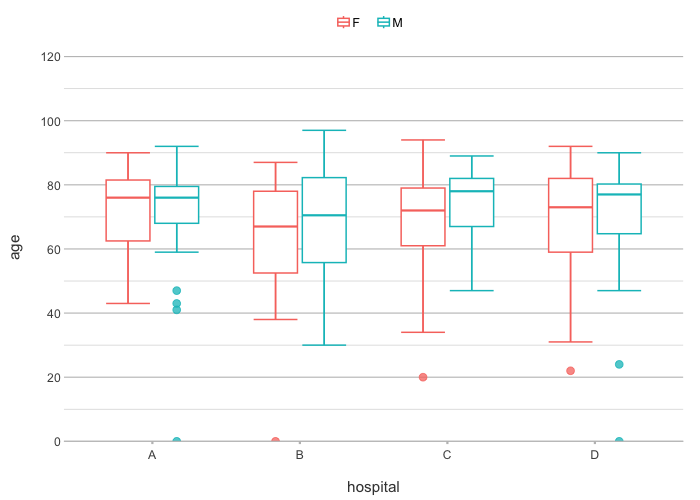
admitted_patients |>
plot2(hospital, age, gender, ward)
#> ℹ Assuming facet.fixed_y = TRUE since the two y scales differ by less than 25%
#> ℹ Assuming facet.repeat_lbls_y = FALSE since y has fixed scales
#> ℹ Using type = "boxplot" since all groups in hospital and gender and ward contain at least three values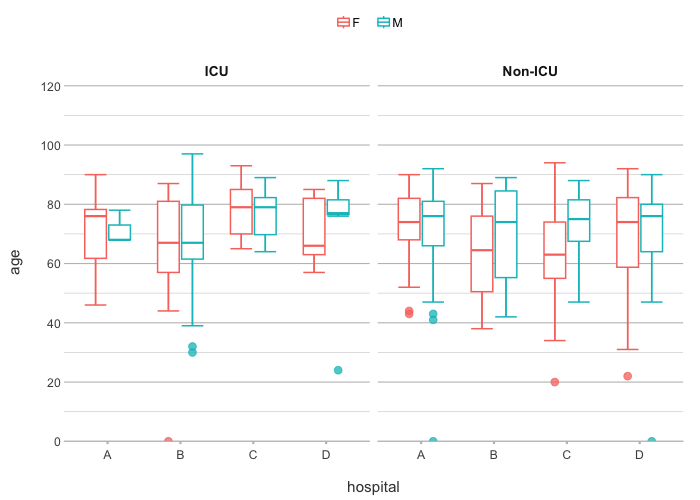
# Use any function for y
admitted_patients |>
plot2(hospital, median(age), gender, ward)
#> ℹ Assuming facet.fixed_y = TRUE since the two y scales differ by less than 25%
#> ℹ Assuming facet.repeat_lbls_y = FALSE since y has fixed scales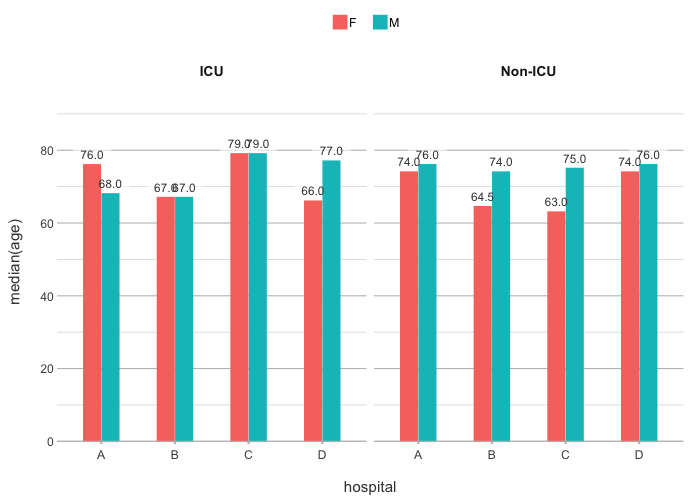
admitted_patients |>
plot2(x = hospital,
y = age,
category = gender,
colour = c("F" = "#3F681C", "M" = "#375E97"),
colour_fill = "#FFBB00",
linewidth = 1.25,
y.age = TRUE)
#> ℹ Using type = "boxplot" since all groups in hospital and gender contain at least three values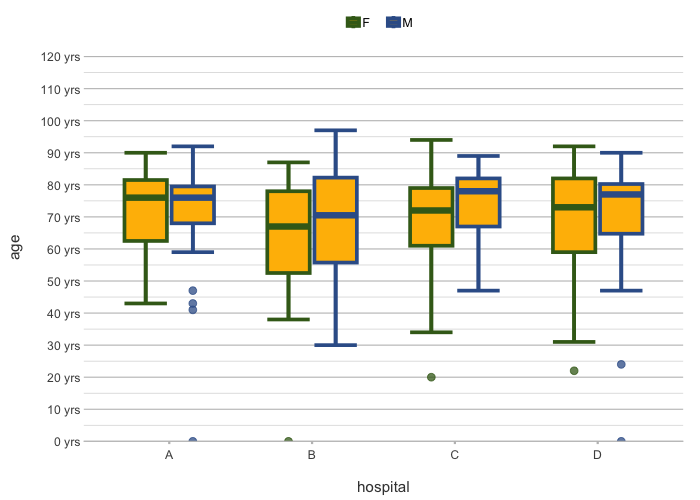
admitted_patients |>
plot2(age, type = "hist")
#> ℹ Assuming smooth = TRUE for type = "histogram"
#> ℹ Using binwidth = 6.4 based on data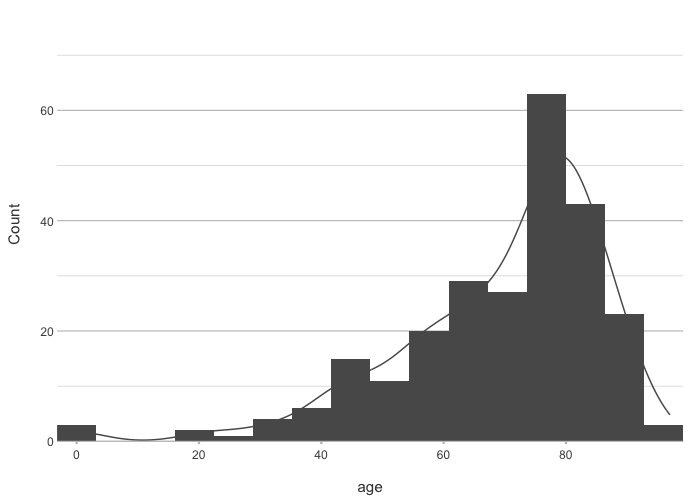
# Titles can include calculations and support {glue}
admitted_patients |>
plot2(age, type = "hist",
title = paste("Based on n =", n_distinct(patient_id), "patients"),
subtitle = paste("Total rows:", n()),
caption = glue::glue("From {n_distinct(hospital)} hospitals"),
x.title = paste("Age ranging from", paste(range(age), collapse = " to ")))
#> ℹ Assuming smooth = TRUE for type = "histogram"
#> ℹ Using binwidth = 6.4 based on data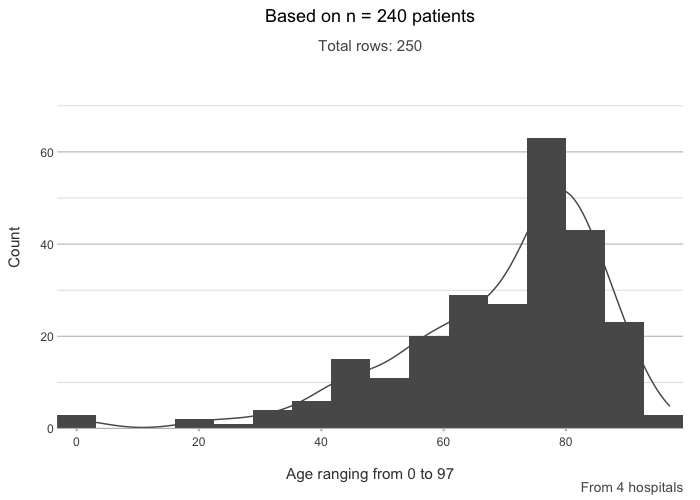
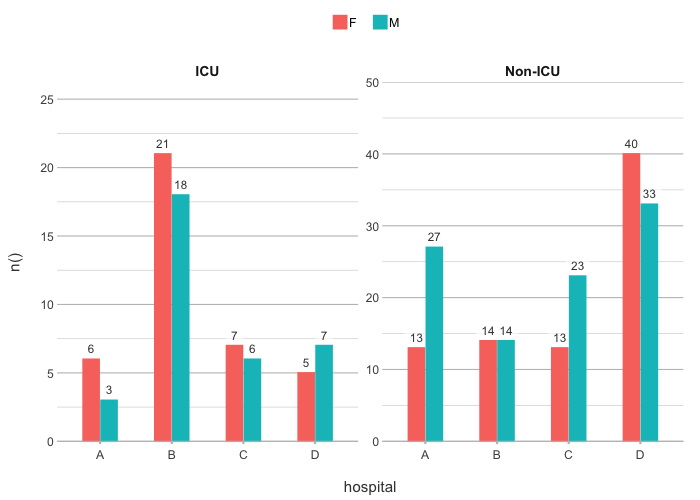
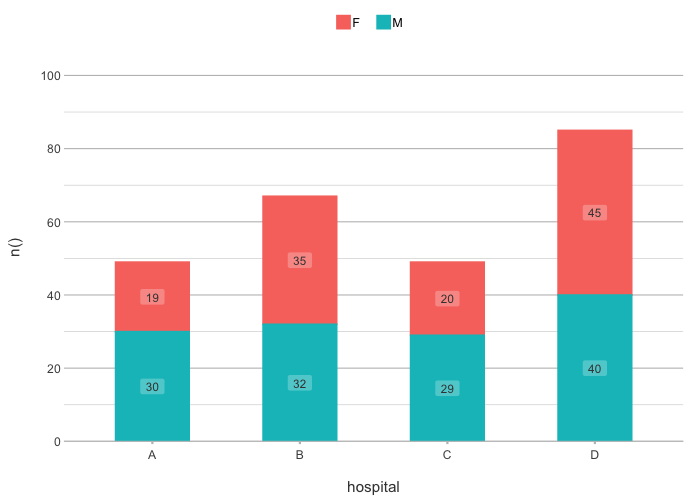
# The default type is column; data labels are automatically
# set for non-continuous types:
admitted_patients |>
plot2(hospital, n(), gender)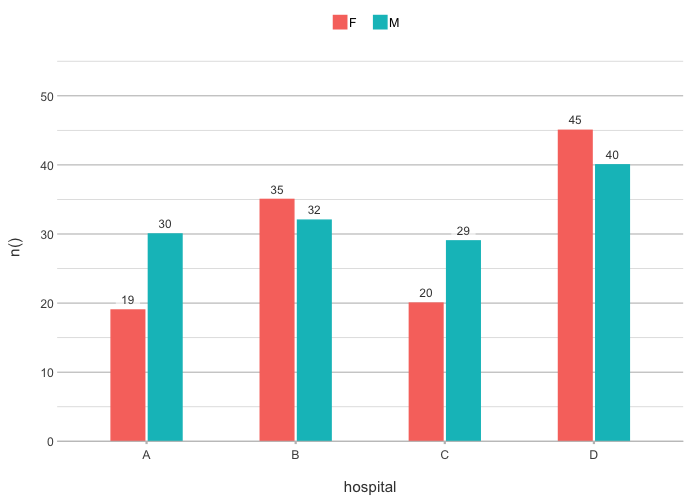
# Two categories may benefit from a dumbbell plot:
admitted_patients |>
plot2(hospital, median(age), gender, type = "dumbbell")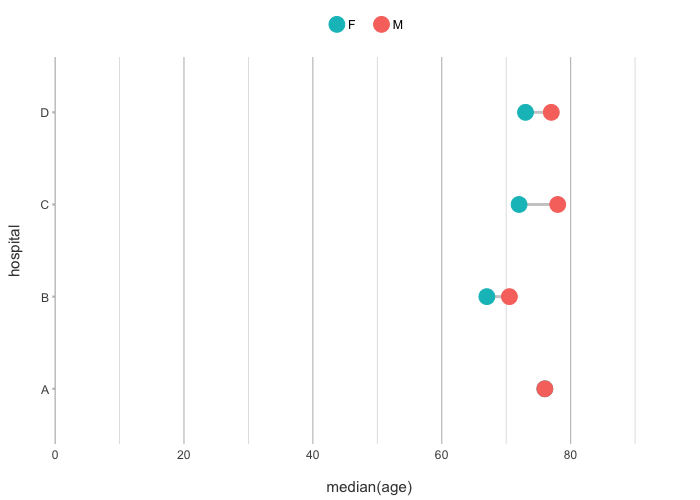
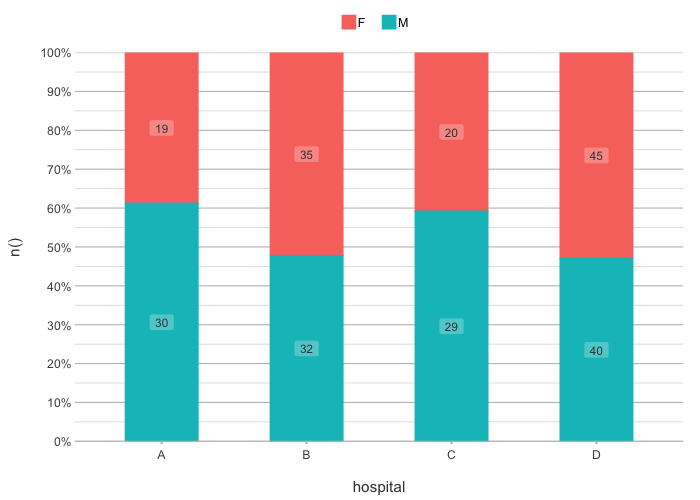
# Sort in any direction:
admitted_patients |>
plot2(hospital, n(), gender,
x.sort = "freq-asc",
stacked = TRUE)
#> ℹ Applying x.sort = "freq-asc" using summarise_function = sum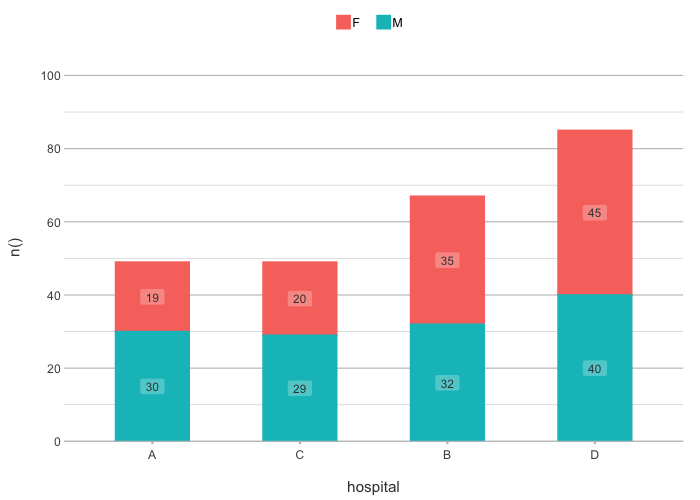
admitted_patients |>
plot2(hospital, n(), gender,
x.sort = c("B", "D", "A"),
category.sort = "alpha-desc",
stacked = TRUE)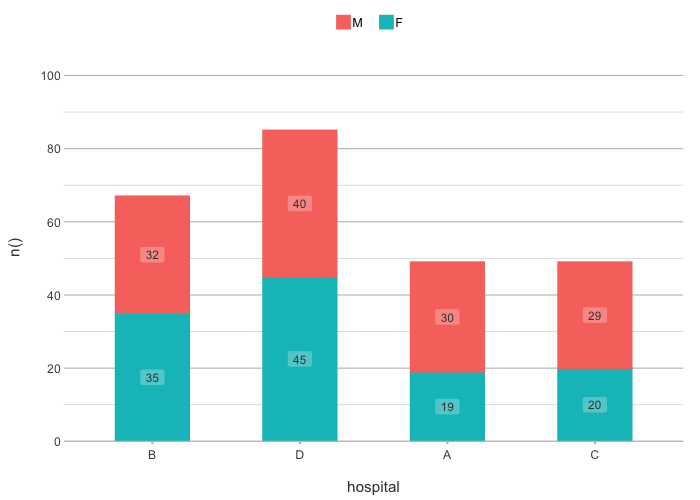
# Support for Sankey plots
Titanic |>
plot2(x = c(Age, Class, Survived),
category = Sex,
type = "sankey")
#> ℹ Assuming sankey.remove_axes = TRUE
#> ! Input class 'table' was transformed using `as.data.frame()`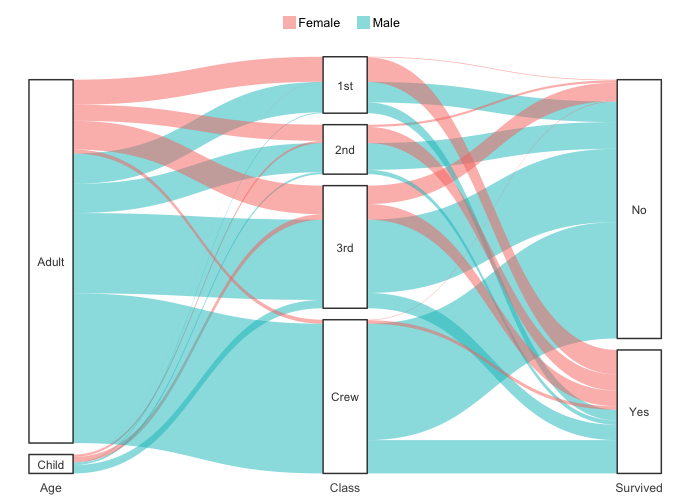
# Matrix support, for example cor()
correlation_matrix <- cor(mtcars)
class(correlation_matrix)
#> [1] "matrix" "array"
head(correlation_matrix)
#> mpg cyl disp hp drat wt qsec vs am gear carb
#> mpg 1.0000000 -0.8521620 -0.8475514 -0.7761684 0.6811719 -0.8676594 0.41868403 0.6640389 0.5998324 0.4802848 -0.5509251
#> cyl -0.8521620 1.0000000 0.9020329 0.8324475 -0.6999381 0.7824958 -0.59124207 -0.8108118 -0.5226070 -0.4926866 0.5269883
#> disp -0.8475514 0.9020329 1.0000000 0.7909486 -0.7102139 0.8879799 -0.43369788 -0.7104159 -0.5912270 -0.5555692 0.3949769
#> hp -0.7761684 0.8324475 0.7909486 1.0000000 -0.4487591 0.6587479 -0.70822339 -0.7230967 -0.2432043 -0.1257043 0.7498125
#> drat 0.6811719 -0.6999381 -0.7102139 -0.4487591 1.0000000 -0.7124406 0.09120476 0.4402785 0.7127111 0.6996101 -0.0907898
#> wt -0.8676594 0.7824958 0.8879799 0.6587479 -0.7124406 1.0000000 -0.17471588 -0.5549157 -0.6924953 -0.5832870 0.4276059
mtcars |>
cor() |>
plot2()
#> ℹ Assuming type = "tile" since the matrix contains identical row and column names
#> ! Omitting printing of 121 datalabels - use datalabels = TRUE to force printing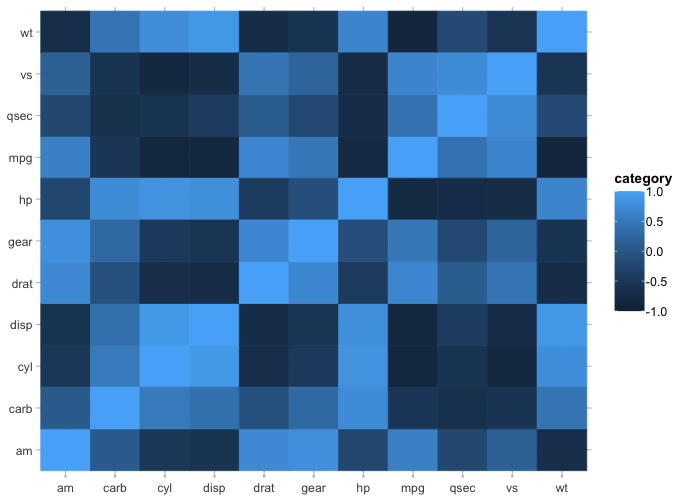
mtcars |>
cor() |>
plot2(colour = c("blue3", "white", "red3"),
datalabels = TRUE,
category.title = "*r*-value",
title = "Correlation matrix")
#> ℹ Assuming type = "tile" since the matrix contains identical row and column names
#> ℹ Using category.midpoint = 0 (the current category scale centre)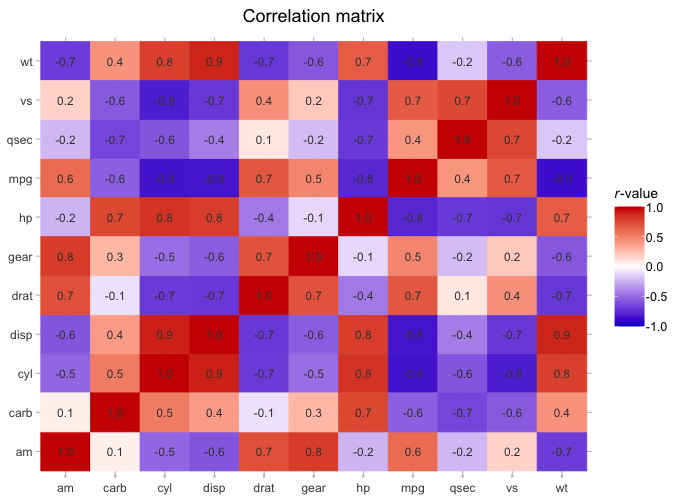
# plot2() supports all S3 extensions available through
# ggplot2::fortify() and broom::augment(), such as regression models:
lm(mpg ~ hp, data = mtcars) |>
plot2(x = mpg ^ -3,
y = hp ^ 2,
smooth = TRUE,
smooth.method = "lm",
smooth.formula = "y ~ log(x)",
title = "Titles and captions support markdown",
subtitle = "Axis titles contain square notation: x^2")
#> ℹ Using type = "point" since both axes are numeric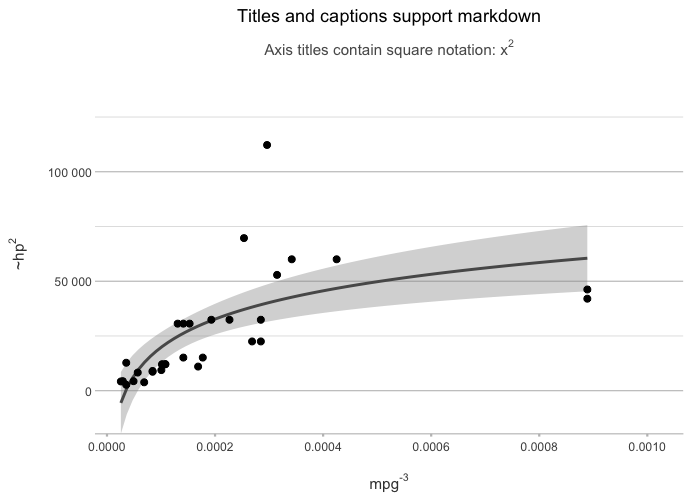
# sf objects, geographic plots using simple features, are also supported
netherlands |>
plot2()
#> ℹ Assuming datalabels.centroid = TRUE. Set to FALSE for a point-on-surface placing of datalabels.
#> ℹ Using category = area_km2
#> ℹ Using datalabels = province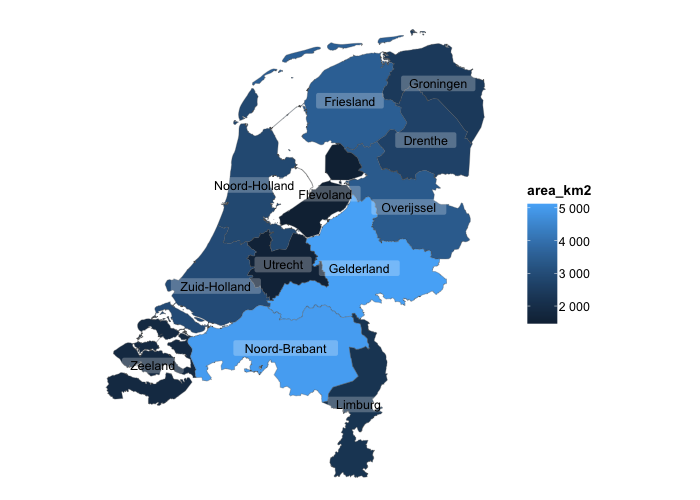
netherlands |>
plot2(colour_fill = "viridis", colour_opacity = 0.75) |>
add_sf(netherlands, colour = "black", colour_fill = NA)
#> ℹ Assuming datalabels.centroid = TRUE. Set to FALSE for a point-on-surface placing of datalabels.
#> ℹ Using category = area_km2
#> ℹ Using datalabels = province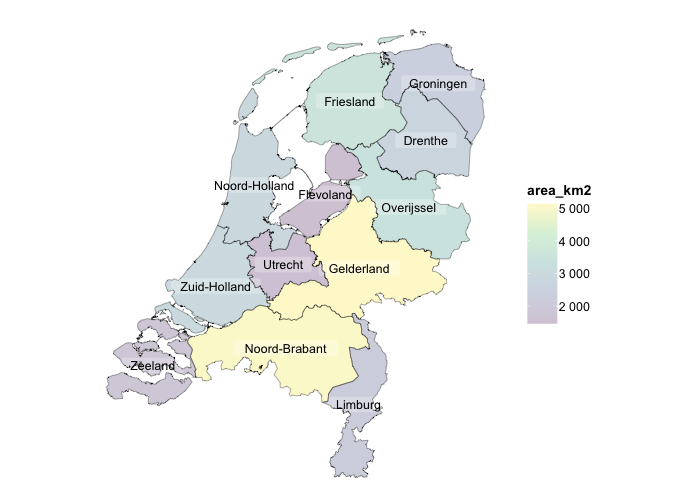
# Support for any system or Google font
mtcars |>
plot2(mpg, hp,
font = "Rock Salt",
title = "This plot uses a Google Font")
#> ℹ Using type = "point" since both axes are numeric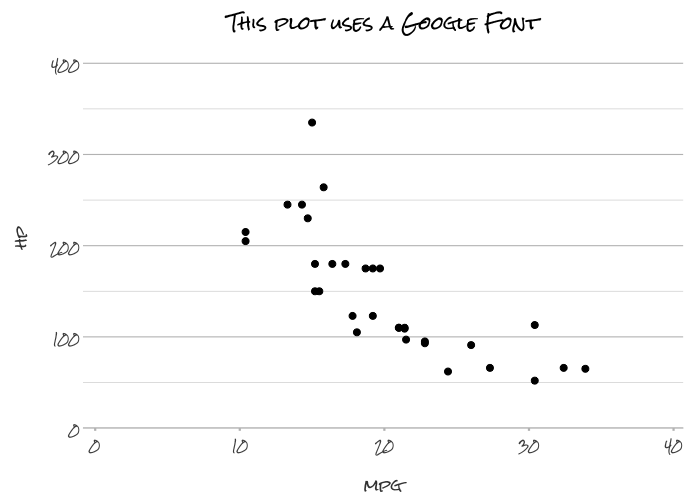
Ways to input arguments
Like plot(), pass x and y in your preferred style:
Shiny app
You can create plots interactively using the built-in Shiny app. All plot2 arguments are available within it. In RStudio, an add-in menu item is also provided.
iris |>
create_interactively()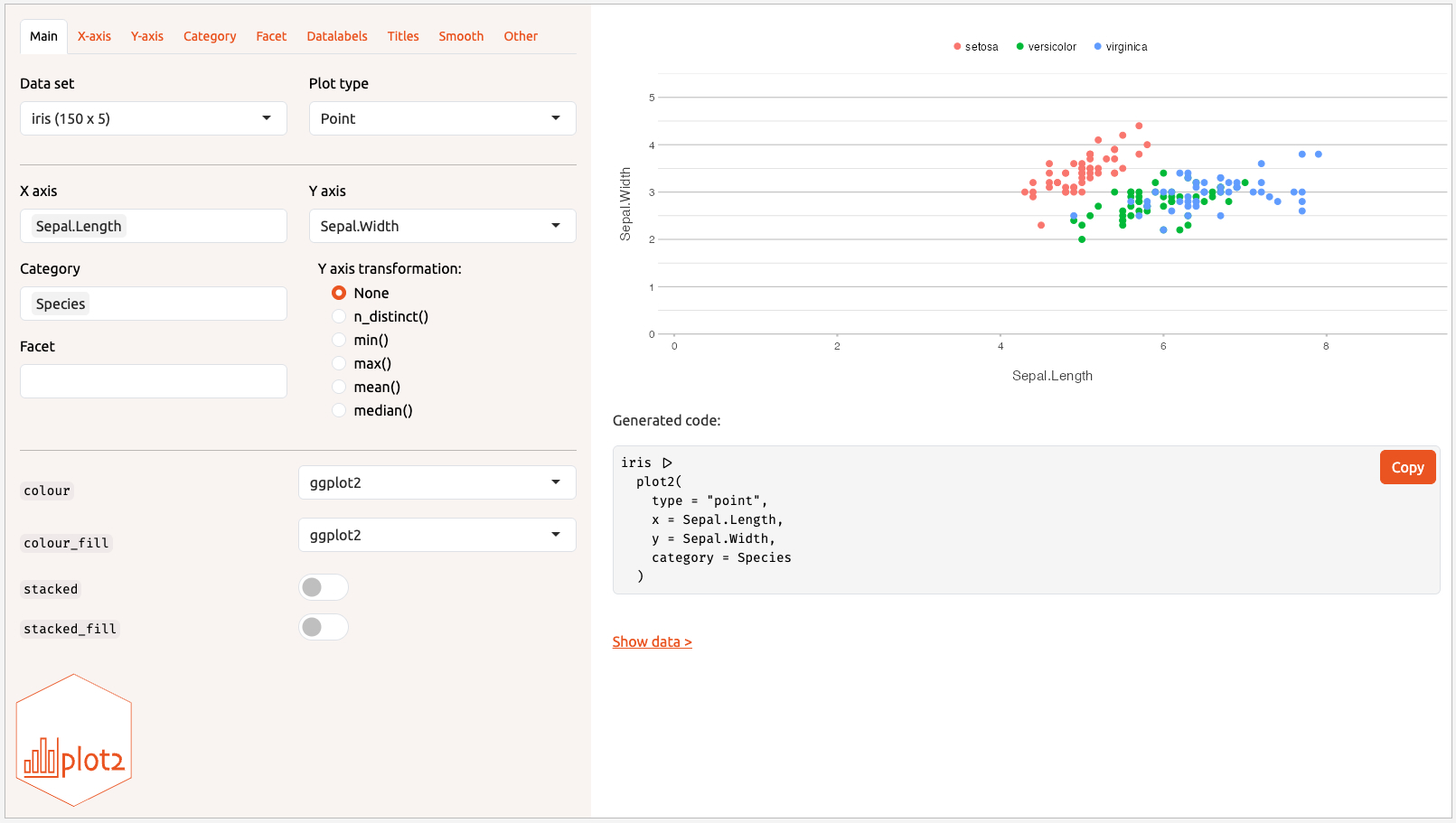
Getting Involved
Contributions to plot2 are welcome, including reporting issues, suggesting features, or submitting pull requests. Familiarity with ggplot2 and the tidyverse is particularly valuable as the package continues to evolve.
Previous Iteration
Although first released here in August 2024, this package builds on years of development and hundreds of Git commits in an earlier iteration available at https://github.com/certe-medical-epidemiology/certeplot2.