Split ages into age groups defined by the split argument. This allows for easier demographic (antimicrobial resistance) analysis.
Usage
age_groups(x, split_at = c(12, 25, 55, 75), na.rm = FALSE)Value
Ordered factor
Details
To split ages, the input for the split_at argument can be:
A numeric vector. A value of e.g.
c(10, 20)will splitxon 0-9, 10-19 and 20+. A value of only50will splitxon 0-49 and 50+. The default is to split on young children (0-11), youth (12-24), young adults (25-54), middle-aged adults (55-74) and elderly (75+).A character:
"children"or"kids", equivalent of:c(0, 1, 2, 4, 6, 13, 18). This will split on 0, 1, 2-3, 4-5, 6-12, 13-17 and 18+."elderly"or"seniors", equivalent of:c(65, 75, 85). This will split on 0-64, 65-74, 75-84, 85+."fives", equivalent of:1:20 * 5. This will split on 0-4, 5-9, ..., 95-99, 100+."tens", equivalent of:1:10 * 10. This will split on 0-9, 10-19, ..., 90-99, 100+.
See also
To determine ages, based on one or more reference dates, use the age() function.
Examples
ages <- c(3, 8, 16, 54, 31, 76, 101, 43, 21)
# split into 0-49 and 50+
age_groups(ages, 50)
#> [1] 0-49 0-49 0-49 50+ 0-49 50+ 50+ 0-49 0-49
#> Levels: 0-49 < 50+
# split into 0-19, 20-49 and 50+
age_groups(ages, c(20, 50))
#> [1] 0-19 0-19 0-19 50+ 20-49 50+ 50+ 20-49 20-49
#> Levels: 0-19 < 20-49 < 50+
# split into groups of ten years
age_groups(ages, 1:10 * 10)
#> [1] 0-9 0-9 10-19 50-59 30-39 70-79 100+ 40-49 20-29
#> 11 Levels: 0-9 < 10-19 < 20-29 < 30-39 < 40-49 < 50-59 < 60-69 < ... < 100+
age_groups(ages, split_at = "tens")
#> [1] 0-9 0-9 10-19 50-59 30-39 70-79 100+ 40-49 20-29
#> 11 Levels: 0-9 < 10-19 < 20-29 < 30-39 < 40-49 < 50-59 < 60-69 < ... < 100+
# split into groups of five years
age_groups(ages, 1:20 * 5)
#> [1] 0-4 5-9 15-19 50-54 30-34 75-79 100+ 40-44 20-24
#> 21 Levels: 0-4 < 5-9 < 10-14 < 15-19 < 20-24 < 25-29 < 30-34 < ... < 100+
age_groups(ages, split_at = "fives")
#> [1] 0-4 5-9 15-19 50-54 30-34 75-79 100+ 40-44 20-24
#> 21 Levels: 0-4 < 5-9 < 10-14 < 15-19 < 20-24 < 25-29 < 30-34 < ... < 100+
# split specifically for children
age_groups(ages, c(1, 2, 4, 6, 13, 18))
#> [1] 2-3 6-12 13-17 18+ 18+ 18+ 18+ 18+ 18+
#> Levels: 0 < 1 < 2-3 < 4-5 < 6-12 < 13-17 < 18+
age_groups(ages, "children")
#> [1] 2-3 6-12 13-17 18+ 18+ 18+ 18+ 18+ 18+
#> Levels: 0 < 1 < 2-3 < 4-5 < 6-12 < 13-17 < 18+
# \donttest{
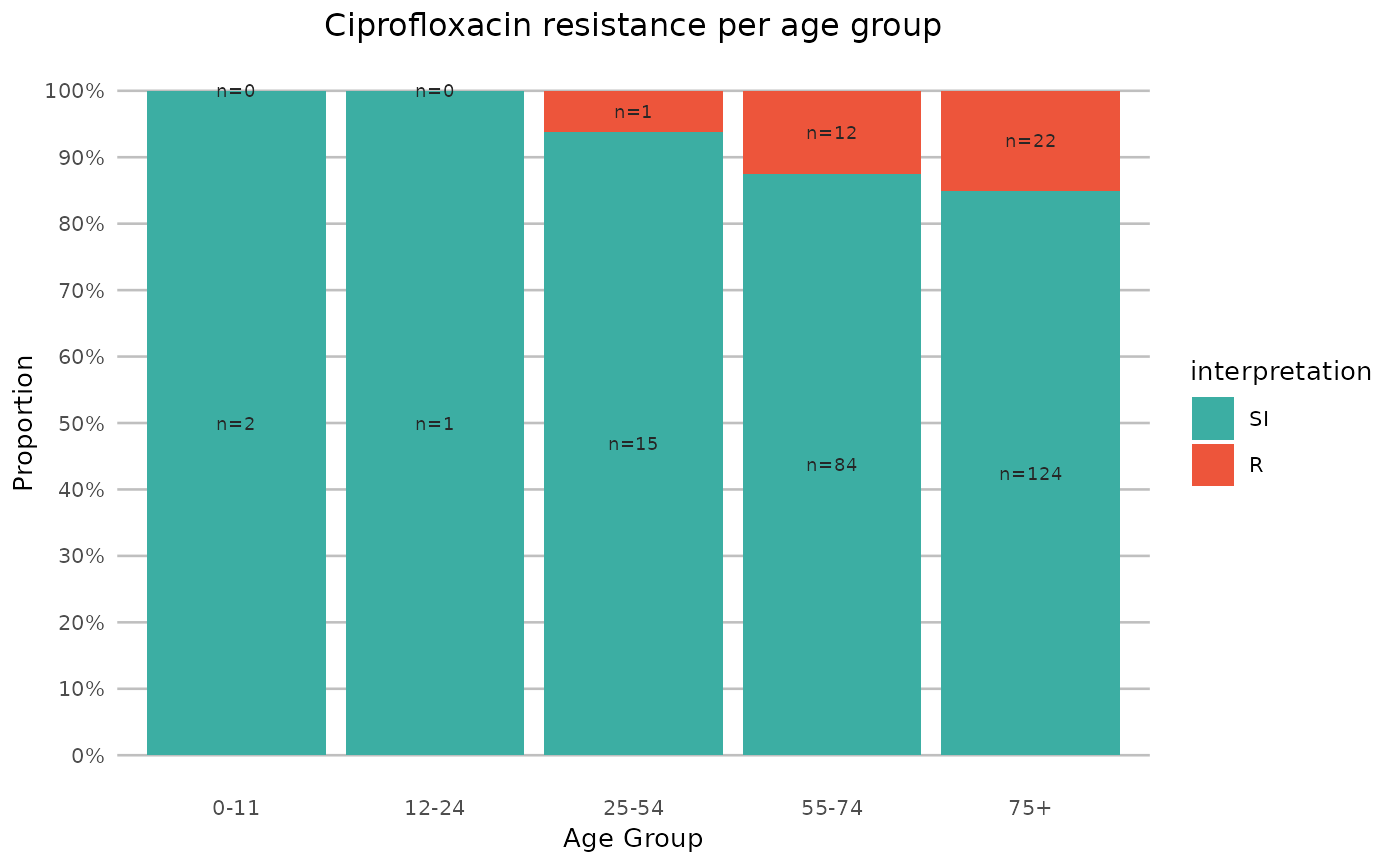
# resistance of ciprofloxacin per age group
if (require("dplyr") && require("ggplot2")) {
example_isolates %>%
filter_first_isolate() %>%
filter(mo == as.mo("Escherichia coli")) %>%
group_by(age_group = age_groups(age)) %>%
select(age_group, CIP) %>%
ggplot_sir(
x = "age_group",
minimum = 0,
x.title = "Age Group",
title = "Ciprofloxacin resistance per age group"
)
}
#> Loading required package: ggplot2
 # }
# }